Automatic Data Aggregation
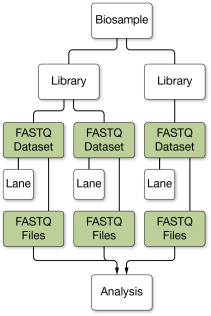
When launching apps that use biosamples as inputs, BaseSpace Sequence Hub automatically aggregates all of the valid (QC Passed) biosample FASTQ data associated with the same default biosamples. You can control which data are used in analyses by setting the QC status of a resource as QC Passed or QC Failed.
NOTE
Aggregated FASTQ data can be produced from multiple libraries, lanes, or flow cells, and can contain data with different read lengths from the same biosample.
To prevent unintended aggregation, biosamples are locked if the biosample was converted from a sample and has data produced by two or more runs. The data cannot be used in analysis until the biosample has been reviewed and unlocked, however you can switch to Classic mode to launch analysis using the original sample data without aggregation. For information about unlocking biosamples to make them available for analysis, see Unlock Biosamples.
NOTE
Classic mode is not supported in all regions.
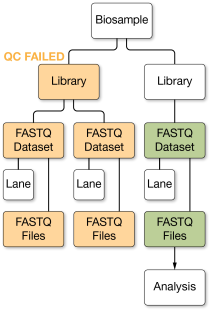
When BaseSpace Sequence Hub collects data for an app launch, it automatically excludes QC-failed lanes, libraries, pools, and any downstream data they produced. For example, if you fail a flow cell lane, all FASTQ data sets produced from that lane are excluded when aggregating data for the biosamples and libraries put on those lanes. If you fail a FASTQ dataset, only that FASTQ dataset is excluded.
If a FASTQ dataset has been copied, BaseSpace Sequence Hub uses the original FASTQ dataset, or the most recent copy if the original is not available. To use a different copy, mark the other copies as QC Failed before starting the analysis.
The following resources can be excluded from data aggregation:
| • | Lanes—Fail lanes using Automatic Lane QC, BaseSpace Sequence Hub API, or manually in BaseSpace Sequence Hub. |
| • | Libraries—Fail libraries using the BaseSpace Sequence Hub API. |
| • | Pools—Fail pools using the BaseSpace Sequence Hub API. |
| • | FASTQ Datasets—Fail FASTQ data sets using the BaseSpace Sequence Hub API, or manually in BaseSpace Sequence Hub. |
NOTE
When using apps that have not been updated to use biosamples or data sets as inputs, BaseSpace Sequence Hub automatically converts the FASTQ data sets into samples before launching the app.
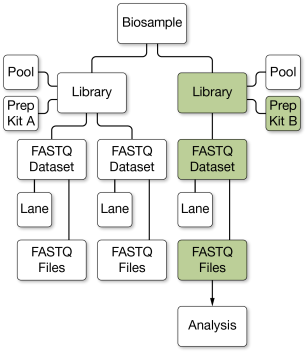
If you specify a library prep kit when selecting a biosample for analysis, the analysis launches using only FASTQ datasets from libraries of the specified prep kit. In the following example, Prep Kit B is selected as input and the FASTQ files from Prep Kit A are excluded.