Biosample Workflow Files
The biosample workflow file is a comma-delimited file that is used to add biosamples in bulk. The file includes a list of biosamples with the following information for each:
| • | Custom metadata |
| • | Library preparation instructions |
| • | Analysis workflows to be launched after sequencing data files have been received |
To make sure that run data are associated with the correct biosample information, upload the biosample workflow file before adding the sample sheet.
Biosample Workflow File Format
Biosample workflow fields are grouped into the following types.
| • | Biosample Data—Lists information about the biosample and default project. See Biosample Data Fields. |
| • | Prep Request Instructions—Instructions about the library preparation for a new or existing biosample. See Prep Request Fields. |
| • | Analysis Workflow—Schedules an analysis workflow for a biosample and lists additional instructions for grouping biosamples and delivering results. See Analysis Workflow Fields. |
| • | Biosample Metadata—Adds custom metadata to the biosample. See Metadata Fields. |
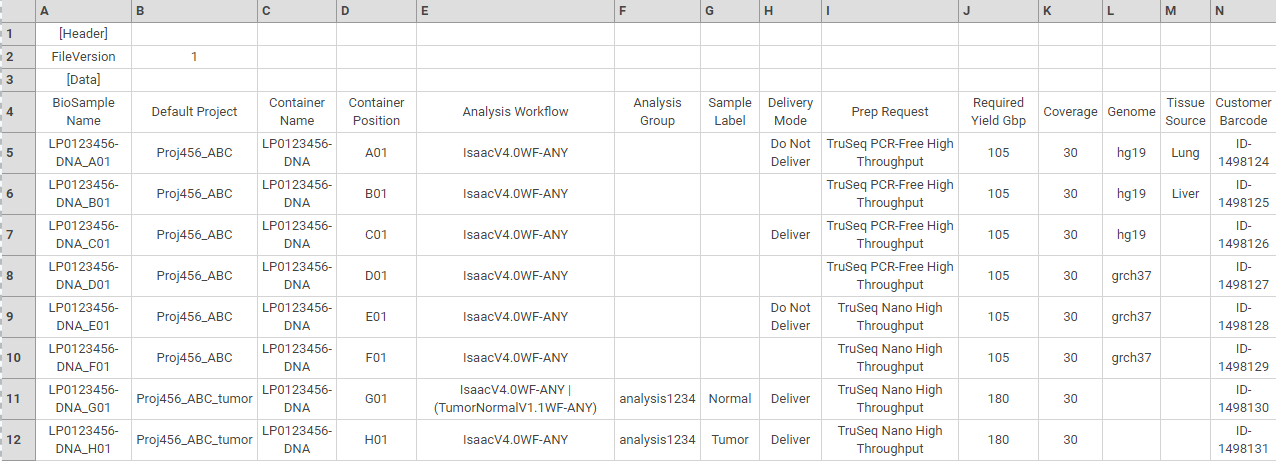
Biosample Data Fields
The biosample data fields define the biosamples you are adding or updating. A biosample and default project combination can only be listed in one row of the manifest. For information about scheduling more than one analysis for a biosample, see Schedule Multiple Analysis Workflows For a Single Biosample.
Analysis Workflow Fields
The analysis workflow fields specify the analysis workflow to launch automatically when sequencing data has been received. For more information, see Analysis Workflows.
| • | For information about scheduling more than one workflow for a biosample, see Schedule Multiple Analysis Workflows For a Single Biosample. |
| • | For information about scheduling an analysis workflow for more than one biosample, see Schedule An Analysis Workflow Using Data From Multiple Biosamples. |
NOTE
Analysis Workflow fields do not overwrite existing scheduled workflows. If you import the same biosample workflow file twice, you duplicate the analysis requests in the file and the analysis is performed a second time.
|
Field |
Requirements |
Description |
||||||
|---|---|---|---|---|---|---|---|---|
|
Analysis Workflow |
The user uploading the biosample workflow must have permission to use the analysis workflow. |
The name of the Analysis Workflow.
|
||||||
|
Analysis Group |
Must be unique within the biosample workflow file. |
Groups multiple biosamples and applies a single analysis workflow to the biosamples in the group. |
||||||
|
Sample Label |
Must be unique within the Analysis Group in the biosample workflow file. |
Labels the analysis workflow inputs that must be distinguished. For example, the Tumor-Normal Analysis Workflow requires that the Isaac Analysis inputs are labeled as 'Normal' and 'Tumor'. Labels are case-sensitive. |
||||||
|
Delivery Mode |
|
Assigns an optional delivery status for an analysis. For example, you can mark R&D or test analyses biosamples as Do Not Deliver. |
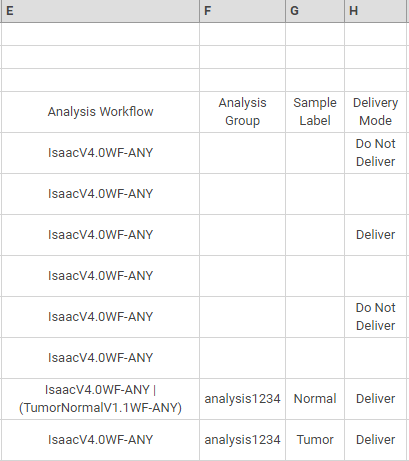
In the following example, columns E—H specify the following information:
| • | The analysis workflows to perform. |
| • | The analysis group and sample labels for a pair of samples analyzed with the Tumor Normal workflow. |
| • | Delivery mode. |
For more information about scheduling Tumor Normal workflows, see Schedule Multiple Analysis Workflows For a Single Biosample and Schedule An Analysis Workflow Using Data From Multiple Biosamples.
Prep Request Fields
You can use the optional prep request fields to do the following:
| • | Specify prep request instructions for a new biosample |
| • | Add a prep request to an existing biosample |
| • | Change the required yield for an existing prep request |
|
Field |
Requirements |
Description |
||||||
|---|---|---|---|---|---|---|---|---|
|
Prep Request |
|
Specifies which library prep to use. |
||||||
|
Required Yield Gbp |
|
Enter a value between 0–1000 to specify the sequencing yield (in Gbp) the lab is required to produce before the planned BaseSpace App can be launched. If the prep request already exists for the biosample, the Required Yield is updated with the new value. |
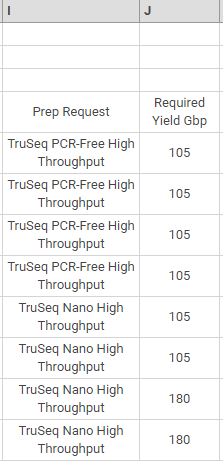
In the following example, columns J—K specify the library prep and required yield.
Metadata Fields
Use metadata fields to add custom information about the biosample. Do not enter patient identifying information in metadata fields.
Biosample metadata is listed on the biosample details page.
NOTE
Metadata can be applied to new biosamples in a biosample workflow file. To add or edit metadata for existing biosamples, use the BaseSpace Sequence Hub API. For more information, see the developer documentation at developer.basespace.illumina.com.
In the following example, columns L—N add genome, tissue source, and customer barcode metadata.
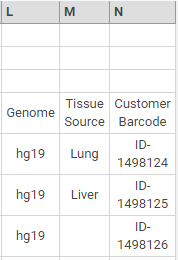
Example Metadata Fields
File Validation
BaseSpace Sequence Hub performs validation on the field requirements and the following general requirements:
| • | Maximum 100 fields |
| • | Field headers must be 4–60 characters |
| • | No special characters in field headers |
| • | Field entries limited to 5000 characters |