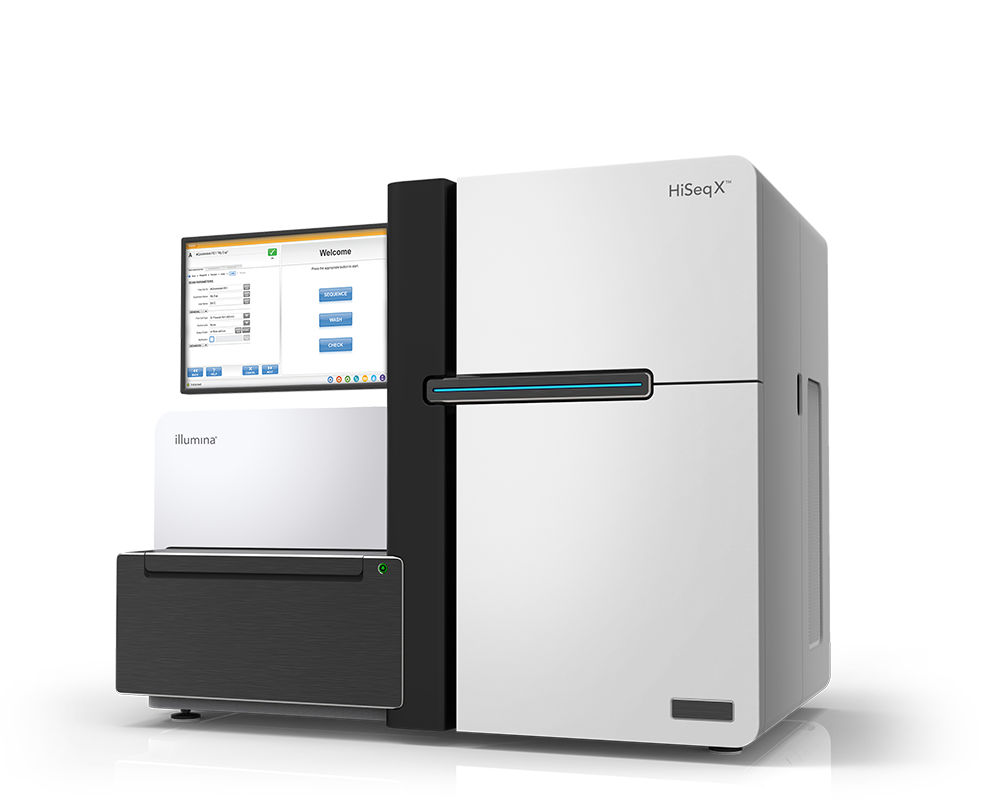
HiSeq X Support
The HiSeq X System is no longer available for purchase. For high-throughput, whole-genome sequencing applications, consider the NovaSeq 6000 System.
Documentation
Software Downloads
Tools
View AllCustom Protocol Selector
Generate end-to-end documentation tailored to your experiment.
Library Prep and Array Kit Selector
Determine the best kit for your project type, starting material, and method or application.
Sequencing Coverage Calculator
Determine reagents and sequencing runs for your desired coverage.
Training
View AllMaximize the effectiveness of your Illumina system, train new employees, or learn the latest techniques and best practices. We offer support webinars, online courses, expert video tips, and instructor-led trainings.